Downloads sampling effort raster files from the Open Science Framework (OSF) based on specified taxonomic groups, descendants, metrics, years, and spatial resolution. The function validates all inputs and retrieves corresponding raster data files. For more information, see this repo and El-Gabbas (2026). Diversity and Distributions (accepted).
Usage
get_sampling_effort(
group = NULL,
descendants = "all",
metric = NULL,
years = "total",
resolution = NULL,
out_dir = getwd(),
conflicts = "skip",
verbose = FALSE
)Arguments
- group
Character. The taxonomic group to download sampling effort data for. Must be one of: "all", "amphibia", "arachnida", "aves", "fungi", "insecta", "mammalia", "mollusca", "reptilia", or "tracheophyta". "all" refers to the overall sampling effort across all groups. Required.
- descendants
A character vector of descendants for the chosen group. Valid descendants vary by group. Defaults to "all" to download data for all descendants combined of the chosen group. See details section below for the list of valid descendants per group. Required.
- metric
A character string specifying the metric to download. Must be either "n_sp" (number of species) or "n_obs" (number of observations). Required.
- years
A numeric vector or character string specifying year(s) to download. Valid range is 1980 to 2025, or "total" (default) for overall efforts. Automatically set to "total" when group is "all". Required for other groups.
- resolution
A numeric value specifying the spatial resolution in kilometers. Must be one of: 1, 5, 10, or 20 km. Required.
- out_dir
A character string specifying the output directory path. Defaults to the current working directory. If the directory does not exist, it will be created.
- conflicts
A character string specifying how to handle existing files. Must be either "skip" (default) or "overwrite".
- verbose
Logical. If
TRUE, prints progress messages during the download process. Defaults toFALSE.
Value
A tibble containing downloaded sampling effort data with columns:
group: The taxonomic groupdescendant: The taxonomic descendantyear: The year of the datametric: The metric type (n_sporn_obs)resolution: The spatial resolution in kmname: The name of the downloaded fileid: The OSF file IDlocal_path: The local file path where the raster was downloadedmeta: Metadata about the downloaded file from OSF
Details
This function downloads sampling effort raster files from the OSF project. This function complements the manuscript "High-resolution, taxon-stratified global sampling effort grids: a reproducible workflow for bias-aware ecological modelling" by providing a programmatic way to access the underlying data used in the study. Please cite the manuscript when using the data retrieved by this function.
Files on the OSF project are organized hierarchically by group, resolution, metric, and descendant-year combinations.
This function retrieves sampling effort rasters based on the specified group
and its descendants. The descendants argument allows users to specify which
descendant groups to download data for. The function checks that the
specified descendants are valid for the chosen group and retrieves the
corresponding raster files from OSF. Valid descendants for each group are as
follows:
all: "all"amphibia: "all", "anura", "caudata", "gymnophiona"arachnida: "all", "amblypygi", "araneae", "holothyrida", "ixodida", "mesostigmata", "opilioacarida", "opiliones", "palpigradi", "pseudoscorpiones", "ricinulei", "sarcoptiformes", "schizomida", "scorpiones", "solifugae", "trombidiformes", "uropygi"aves: "all", "accipitriformes", "anseriformes", "apodiformes", "apterygiformes", "bucerotiformes", "caprimulgiformes", "cariamiformes", "casuariiformes", "charadriiformes", "ciconiiformes", "coliiformes", "columbiformes", "coraciiformes", "cuculiformes", "eurypygiformes", "falconiformes", "galliformes", "gaviiformes", "gruiformes", "leptosomiformes", "mesitornithiformes", "musophagiformes", "nyctibiiformes", "opisthocomiformes", "otidiformes", "passeriformes", "pelecaniformes", "phaethontiformes", "phoenicopteriformes", "piciformes", "podicipediformes", "procellariiformes", "psittaciformes", "pteroclidiformes", "rheiformes", "sphenisciformes", "steatornithiformes", "strigiformes", "struthioniformes", "suliformes", "tinamiformes", "trogoniformes"fungi: "all", "ascomycota", "basidiomycota", "blastocladiomycota", "chytridiomycota", "entomophthoromycota", "glomeromycota", "mucoromycota", "neocallimastigomycota", "sanchytriomycota", "zoopagomycota", "zygomycota"insecta: "all", "archaeognatha", "blattodea", "cnemidolestodea", "coleoptera", "dermaptera", "diptera", "embioptera", "ephemeroptera", "grylloblattodea", "hemiptera", "hymenoptera", "lepidoptera", "mantodea", "mantophasmatodea", "mecoptera", "megaloptera", "neuroptera", "odonata", "orthoptera", "palaeodictyoptera", "phasmida", "plecoptera", "protorthoptera", "psocodea", "raphidioptera", "siphonaptera", "strepsiptera", "thysanoptera", "trichoptera", "zoraptera", "zygentoma"mammalia: "all", "afrosoricida", "artiodactyla", "carnivora", "cetacea", "chiroptera", "cingulata", "dasyuromorphia", "dermoptera", "didelphimorphia", "diprotodontia", "erinaceomorpha", "hyracoidea", "lagomorpha", "macroscelidea", "microbiotheria", "monotremata", "notoryctemorphia", "paucituberculata", "peramelemorphia", "perissodactyla", "pholidota", "pilosa", "primates", "proboscidea", "rodentia", "scandentia", "sirenia", "soricomorpha", "tubulidentata"mollusca: "all", "bivalvia", "caudofoveata", "cephalopoda", "cricoconarida", "gastropoda", "monoplacophora", "polyplacophora", "rostroconchia", "scaphopoda", "solenogastres"reptilia: "all", "crocodylia", "sphenodontia", "squamata", "testudines"tracheophyta: "all", "cycadopsida", "ginkgoopsida", "gnetopsida", "liliopsida", "lycopodiopsida", "magnoliopsida", "pinopsida", "polypodiopsida"
Examples
require(terra)
# Occurrence count for birds at 20 km resolution
efforts_birds_all <- get_sampling_effort(
group = "aves", metric = "n_obs", resolution = 20)
dplyr::glimpse(efforts_birds_all)
#> Rows: 1
#> Columns: 9
#> $ group <chr> "aves"
#> $ descendant <chr> "all"
#> $ year <chr> "total"
#> $ metric <chr> "n_obs"
#> $ resolution <dbl> 20
#> $ name <chr> "n_obs_Aves_res_20.tif"
#> $ id <chr> "69144be425b8c888ea3ee2b8"
#> $ local_path <chr> "./n_obs_Aves_res_20.tif"
#> $ meta <list> [[<NULL>, <NULL>, "n_obs_Aves_res_20.tif", "file", "/69144b…
efforts_birds_all_r <- terra::rast(efforts_birds_all$local_path)
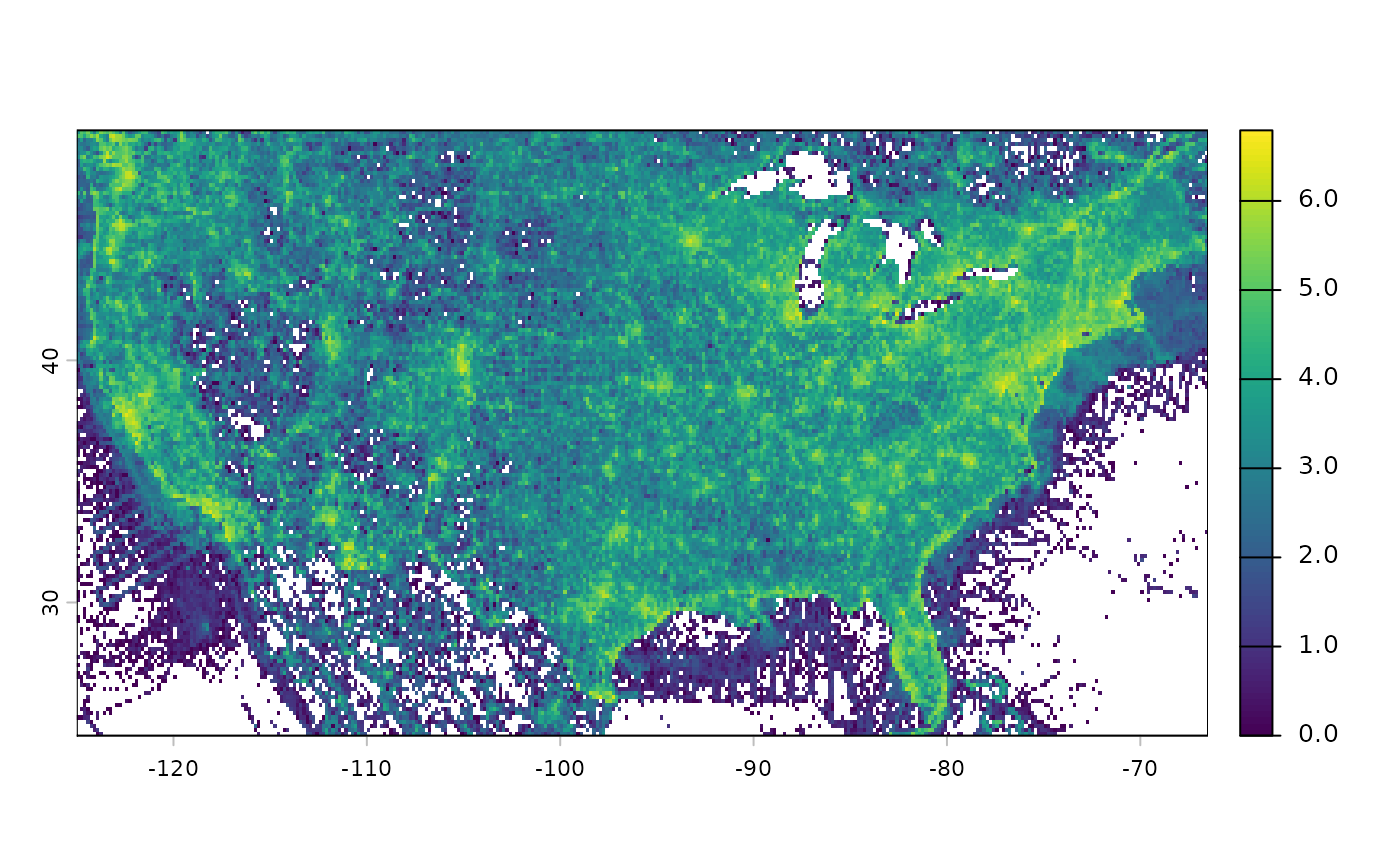
# Plot at log10 scale
terra::classify(efforts_birds_all_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
log10() %>%
plot()
 # |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
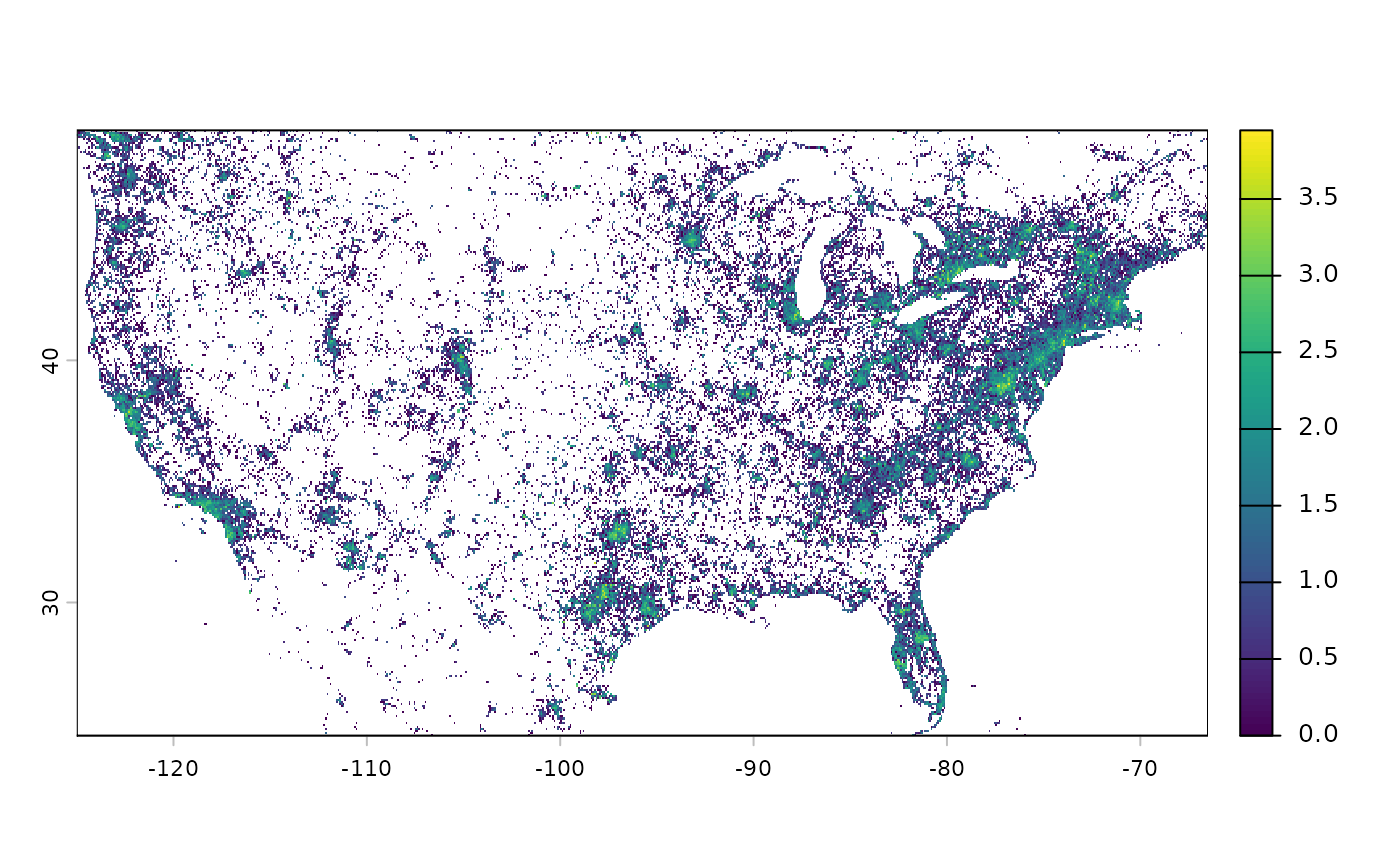
# Occurrence count for insecta at 10 km resolution in 2020
efforts_insecta_2020 <- get_sampling_effort(
group = "insecta", descendants = "all",
metric = "n_obs", years = 2020, resolution = 10)
efforts_insecta_2020_r <- terra::rast(efforts_insecta_2020$local_path)
# Plot at log10 scale
terra::classify(efforts_insecta_2020_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
log10() %>%
plot()
# |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
# Occurrence count for insecta at 10 km resolution in 2020
efforts_insecta_2020 <- get_sampling_effort(
group = "insecta", descendants = "all",
metric = "n_obs", years = 2020, resolution = 10)
efforts_insecta_2020_r <- terra::rast(efforts_insecta_2020$local_path)
# Plot at log10 scale
terra::classify(efforts_insecta_2020_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
log10() %>%
plot()
 # |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
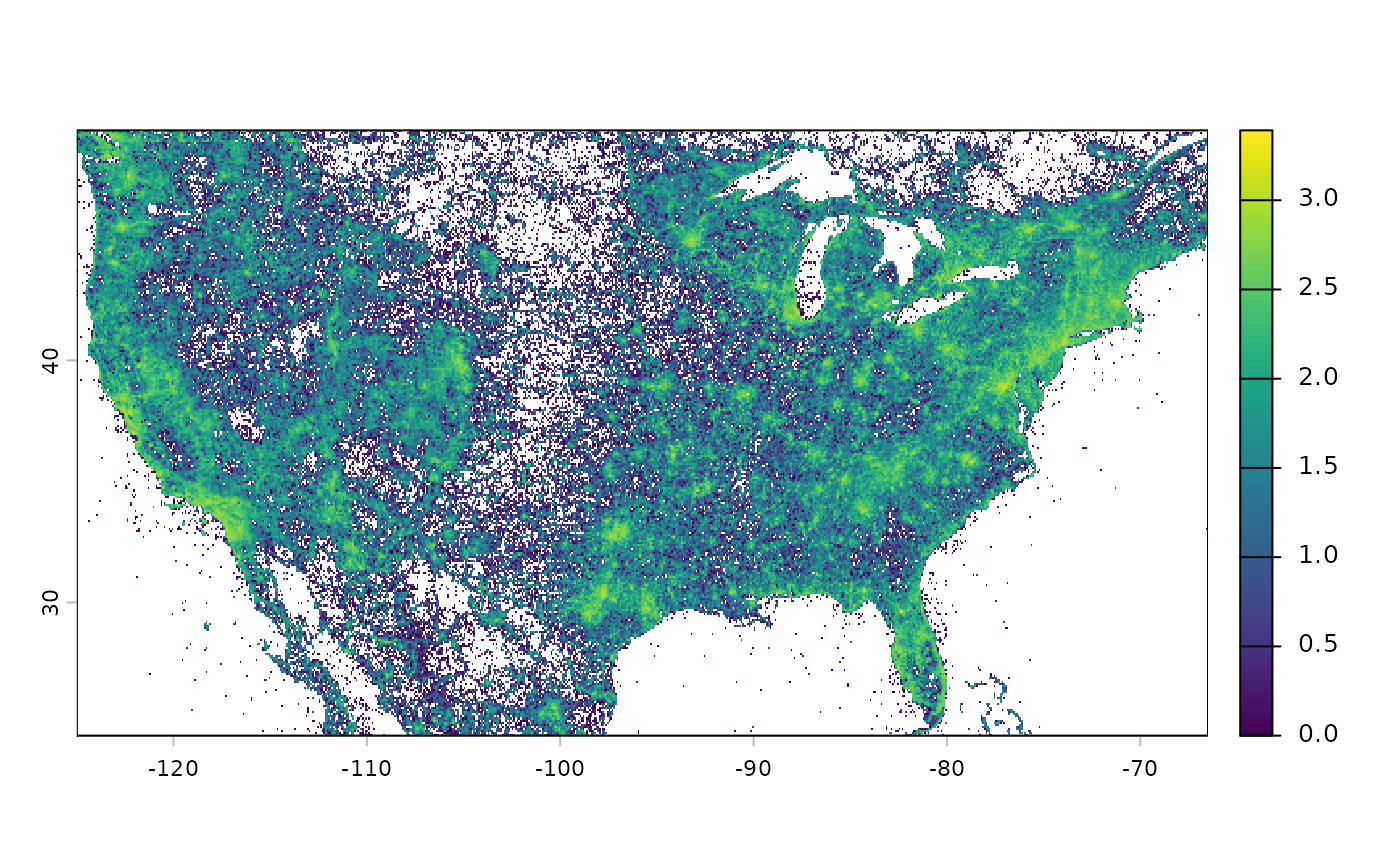
# Species count for vascular plants at 10 km
efforts_plants_2020 <- get_sampling_effort(
group = "tracheophyta", descendants = "all",
metric = "n_sp", resolution = 10)
efforts_plants_2020_r <- terra::rast(efforts_plants_2020$local_path)
# Plot at log10 scale
terra::classify(efforts_plants_2020_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
log10() %>%
plot()
# |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
# Species count for vascular plants at 10 km
efforts_plants_2020 <- get_sampling_effort(
group = "tracheophyta", descendants = "all",
metric = "n_sp", resolution = 10)
efforts_plants_2020_r <- terra::rast(efforts_plants_2020$local_path)
# Plot at log10 scale
terra::classify(efforts_plants_2020_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
log10() %>%
plot()
 # |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
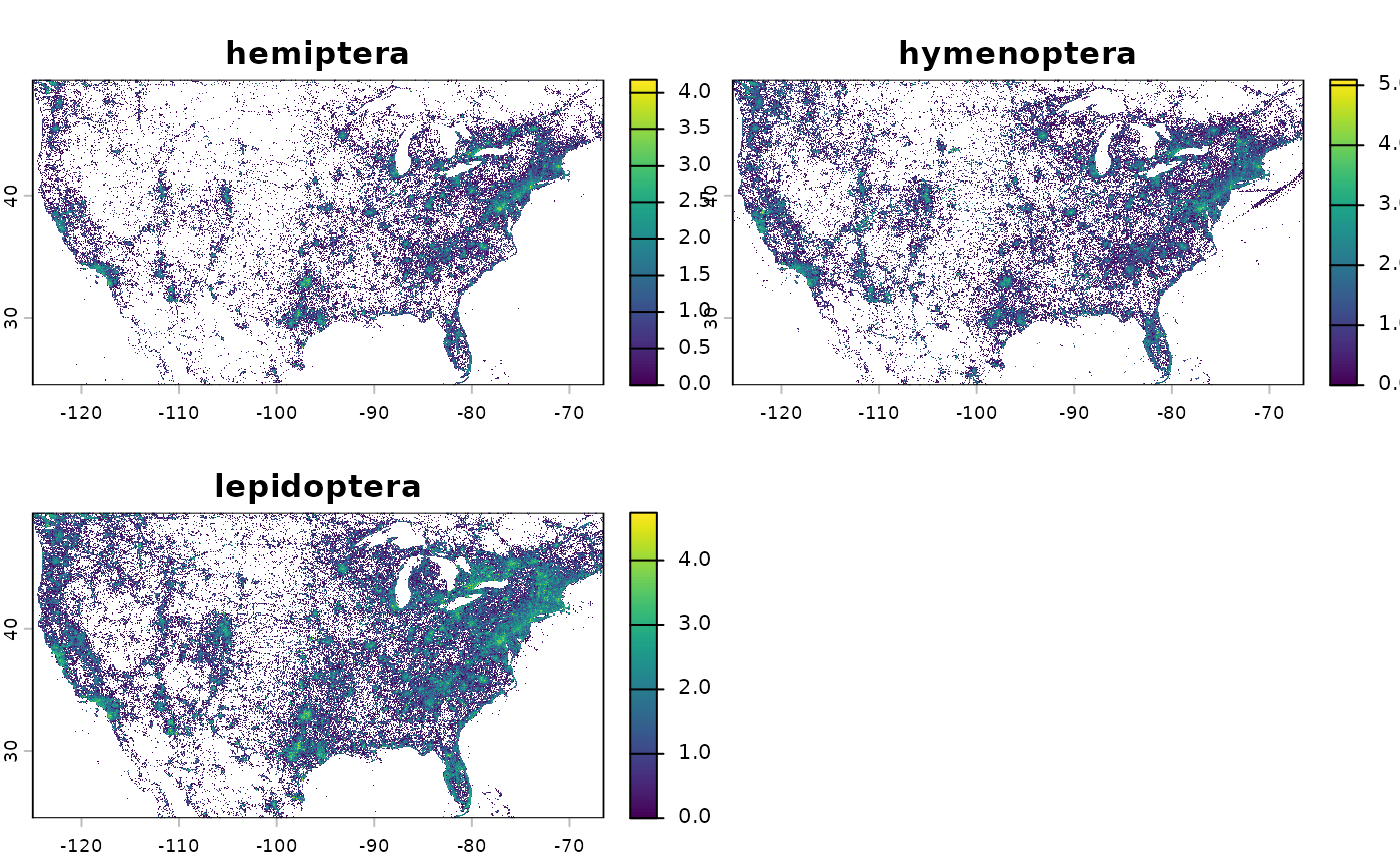
# Occurrence count for selected insect groups (descendants) at 10 km
efforts_insects <- get_sampling_effort(
group = "insecta",
descendants = c("hemiptera", "hymenoptera", "lepidoptera"),
metric = "n_obs", resolution = 10)
efforts_insects
#> # A tibble: 3 × 9
#> group descendant year metric resolution name id local_path meta
#> <chr> <chr> <chr> <chr> <dbl> <chr> <chr> <chr> <list>
#> 1 insecta hemiptera total n_obs 10 n_ob… 6916… ./n_obs_H… <named list>
#> 2 insecta hymenopte… total n_obs 10 n_ob… 6916… ./n_obs_H… <named list>
#> 3 insecta lepidopte… total n_obs 10 n_ob… 6916… ./n_obs_L… <named list>
dplyr::glimpse(efforts_insects)
#> Rows: 3
#> Columns: 9
#> $ group <chr> "insecta", "insecta", "insecta"
#> $ descendant <chr> "hemiptera", "hymenoptera", "lepidoptera"
#> $ year <chr> "total", "total", "total"
#> $ metric <chr> "n_obs", "n_obs", "n_obs"
#> $ resolution <dbl> 10, 10, 10
#> $ name <chr> "n_obs_Hemiptera_total_res_10.tif", "n_obs_Hymenoptera_tota…
#> $ id <chr> "691627285d3006de940c2892", "6916281e843c090b4dfdc3f1", "69…
#> $ local_path <chr> "./n_obs_Hemiptera_total_res_10.tif", "./n_obs_Hymenoptera_…
#> $ meta <list> [[<NULL>, <NULL>, "n_obs_Hemiptera_total_res_10.tif", "file…
efforts_insects_r <- terra::rast(efforts_insects$local_path)
efforts_insects_r
#> class : SpatRaster
#> size : 2160, 4320, 3 (nrow, ncol, nlyr)
#> resolution : 0.08333333, 0.08333333 (x, y)
#> extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
#> coord. ref. : lon/lat WGS 84 (EPSG:4326)
#> sources : n_obs_Hemiptera_total_res_10.tif
#> n_obs_Hymenoptera_total_res_10.tif
#> n_obs_Lepidoptera_total_res_10.tif
#> names : n_obs_Hemi~tal_res_10, n_obs_Hyme~tal_res_10, n_obs_Lepi~tal_res_10
#> min values : 0, 0, 0
#> max values : 92257, 270776, 324427
# Plot at log10 scale
terra::classify(efforts_insects_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
stats::setNames(c("hemiptera", "hymenoptera", "lepidoptera")) %>%
log10() %>%
plot()
# |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
# Occurrence count for selected insect groups (descendants) at 10 km
efforts_insects <- get_sampling_effort(
group = "insecta",
descendants = c("hemiptera", "hymenoptera", "lepidoptera"),
metric = "n_obs", resolution = 10)
efforts_insects
#> # A tibble: 3 × 9
#> group descendant year metric resolution name id local_path meta
#> <chr> <chr> <chr> <chr> <dbl> <chr> <chr> <chr> <list>
#> 1 insecta hemiptera total n_obs 10 n_ob… 6916… ./n_obs_H… <named list>
#> 2 insecta hymenopte… total n_obs 10 n_ob… 6916… ./n_obs_H… <named list>
#> 3 insecta lepidopte… total n_obs 10 n_ob… 6916… ./n_obs_L… <named list>
dplyr::glimpse(efforts_insects)
#> Rows: 3
#> Columns: 9
#> $ group <chr> "insecta", "insecta", "insecta"
#> $ descendant <chr> "hemiptera", "hymenoptera", "lepidoptera"
#> $ year <chr> "total", "total", "total"
#> $ metric <chr> "n_obs", "n_obs", "n_obs"
#> $ resolution <dbl> 10, 10, 10
#> $ name <chr> "n_obs_Hemiptera_total_res_10.tif", "n_obs_Hymenoptera_tota…
#> $ id <chr> "691627285d3006de940c2892", "6916281e843c090b4dfdc3f1", "69…
#> $ local_path <chr> "./n_obs_Hemiptera_total_res_10.tif", "./n_obs_Hymenoptera_…
#> $ meta <list> [[<NULL>, <NULL>, "n_obs_Hemiptera_total_res_10.tif", "file…
efforts_insects_r <- terra::rast(efforts_insects$local_path)
efforts_insects_r
#> class : SpatRaster
#> size : 2160, 4320, 3 (nrow, ncol, nlyr)
#> resolution : 0.08333333, 0.08333333 (x, y)
#> extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
#> coord. ref. : lon/lat WGS 84 (EPSG:4326)
#> sources : n_obs_Hemiptera_total_res_10.tif
#> n_obs_Hymenoptera_total_res_10.tif
#> n_obs_Lepidoptera_total_res_10.tif
#> names : n_obs_Hemi~tal_res_10, n_obs_Hyme~tal_res_10, n_obs_Lepi~tal_res_10
#> min values : 0, 0, 0
#> max values : 92257, 270776, 324427
# Plot at log10 scale
terra::classify(efforts_insects_r, cbind(0, NA)) %>%
terra::crop(terra::ext(-125, -66.5, 24.5, 49.5)) %>%
stats::setNames(c("hemiptera", "hymenoptera", "lepidoptera")) %>%
log10() %>%
plot()